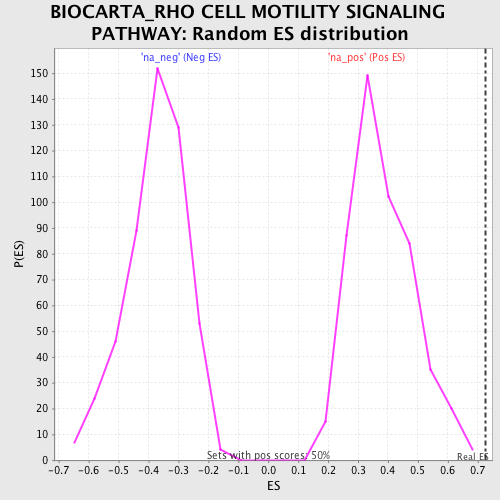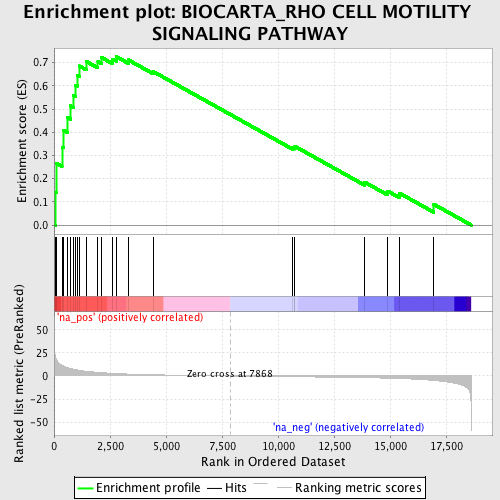
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_DMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | BIOCARTA_RHO CELL MOTILITY SIGNALING PATHWAY |
| Enrichment Score (ES) | 0.72523063 |
| Normalized Enrichment Score (NES) | 1.901957 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.053037096 |
| FWER p-Value | 0.173 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MYLK | 71 | 20.288 | 0.1402 | Yes | ||
| 2 | ARHGEF5 | 97 | 18.082 | 0.2672 | Yes | ||
| 3 | VAV1 | 354 | 11.615 | 0.3359 | Yes | ||
| 4 | ARFGAP3 | 410 | 10.819 | 0.4097 | Yes | ||
| 5 | RALBP1 | 597 | 9.033 | 0.4638 | Yes | ||
| 6 | VCL | 736 | 8.130 | 0.5141 | Yes | ||
| 7 | NCF2 | 852 | 7.431 | 0.5607 | Yes | ||
| 8 | ARHGAP4 | 967 | 6.888 | 0.6034 | Yes | ||
| 9 | ARHGEF1 | 1043 | 6.559 | 0.6459 | Yes | ||
| 10 | PIP5K1A | 1125 | 6.253 | 0.6860 | Yes | ||
| 11 | PIP5K1B | 1448 | 5.095 | 0.7048 | Yes | ||
| 12 | GSN | 1957 | 3.905 | 0.7052 | Yes | ||
| 13 | TLN1 | 2105 | 3.630 | 0.7231 | Yes | ||
| 14 | MYL2 | 2614 | 2.843 | 0.7159 | Yes | ||
| 15 | ARHGAP6 | 2790 | 2.639 | 0.7252 | Yes | ||
| 16 | ROCK1 | 3311 | 2.094 | 0.7121 | No | ||
| 17 | LIMK1 | 4422 | 1.338 | 0.6619 | No | ||
| 18 | CACNA1A | 10653 | -0.811 | 0.3326 | No | ||
| 19 | RHOA | 10740 | -0.836 | 0.3339 | No | ||
| 20 | ARFGAP1 | 10750 | -0.838 | 0.3394 | No | ||
| 21 | ARHGAP5 | 13857 | -1.836 | 0.1854 | No | ||
| 22 | PFN1 | 14888 | -2.358 | 0.1467 | No | ||
| 23 | ARHGAP1 | 15417 | -2.732 | 0.1377 | No | ||
| 24 | CFL1 | 16942 | -4.834 | 0.0900 | No |